Lab Web Sites
 |
Altman Lab The Helix Group at Stanford is directed by Russ Altman. |
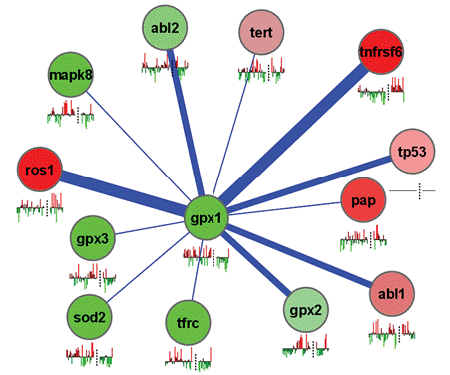 |
Ashley Lab The Ashley lab is focused on the application of genomics to medicine. |
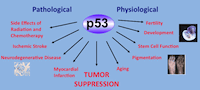 |
Attardi Lab The overarching goal of our research is to better define the mechanisms by which the p53 protein promotes different responses in different settings. |
 |
Baker Lab Cellular differentiation is governed by dynamic changes occurring in the genome. |
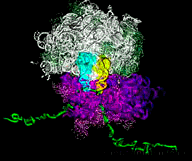 |
Barna Lab We study how the genome is translated into morphology through a ribosome code and single cell imaging of tissue patterning. |
 |
Bhatt Lab The Bhatt lab applies modern genetic tools to deconvolute how the microbiome is intertwined with states of health and disease. |
 |
Brunet Lab Our laboratory studies the molecular mechanisms of aging and longevity. |
 |
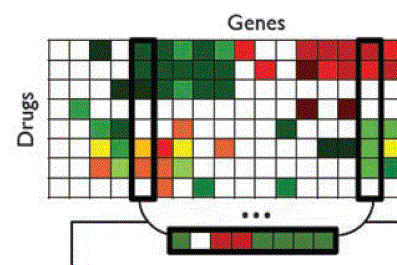
Bustamante Lab Analyzing genome wide patterns of variation to address fundamental questions in biology, anthropology, and medicine using computational biology, mathematical genetics, and evolutionary genomics. |
 |
Butte Lab Our lab aims to address fundamental and therapeutic questions in immunology by developing and using tools from soft lithography and advanced microscopy to visualize and manipulate cells. |
 |
Calos Lab The Calos Lab is interested in developing novel gene and cell therapy approaches to address human diseases. |
 |
Cherry Lab Innovation in literature curation, dataset validation and ontologies enhance experimental results. |
 |
Cohen Lab We study RNA decay, microbial antibiotic resistance, and mechanisms that regulate transcription elongation through genes containing expanded regions of trinucleotide repeats. |
 |
Curtis Lab We aim to characterize the evolutionary dynamics of tumor progression and the genotype-phenotype map in cancer by leveraging both experimental and computational approaches. |
 |
Davis Lab Our center develops new technologies to address important biological questions that otherwise would not be feasible. |
 |
Fire Lab The Fire Lab studies the mechanisms by which cells and organisms respond to genetic change. |
 |
Ford Lab The major focus of this laboratory is to explore the mammalian genetic determinants of the inducible response and cellular sensitivity. |
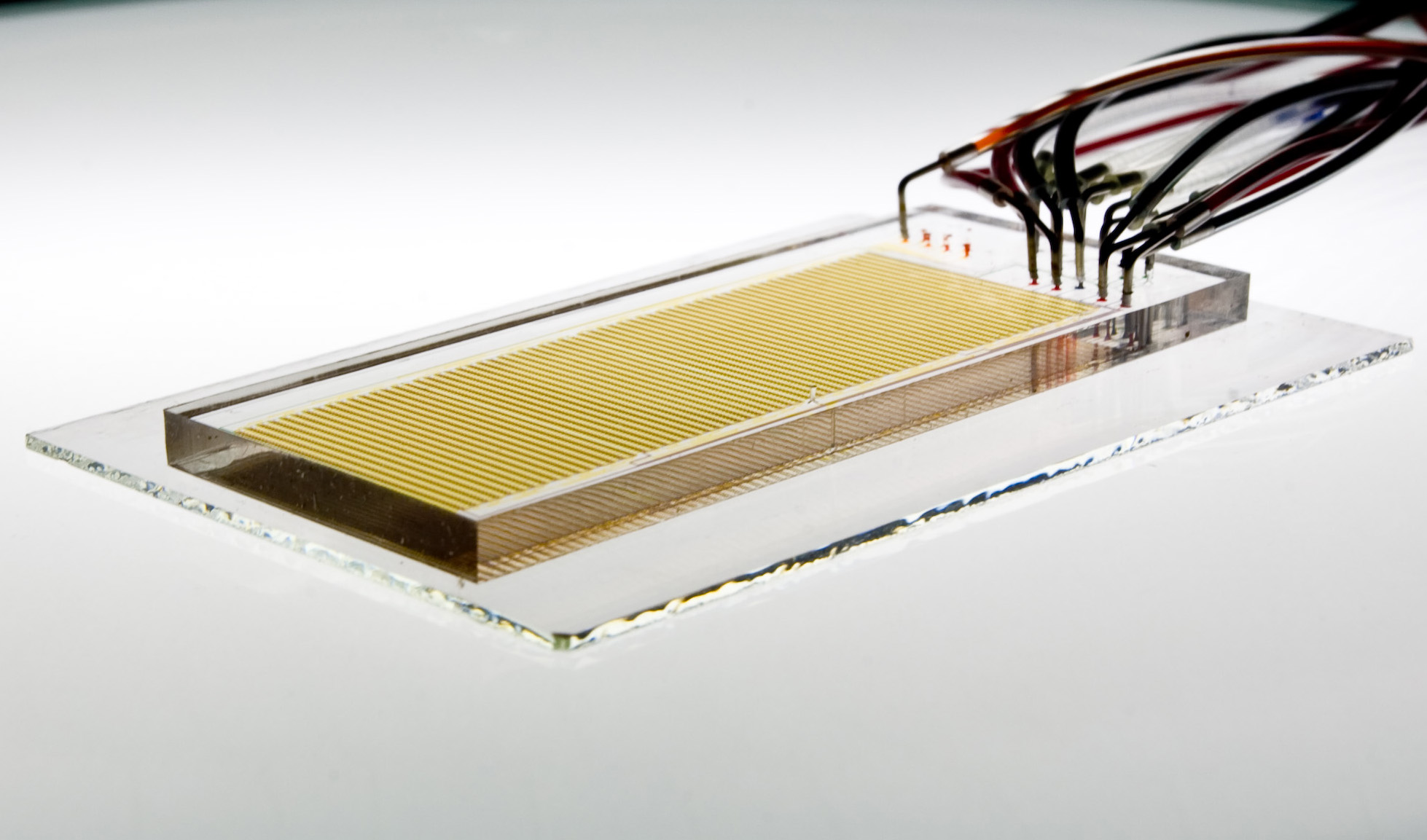 |
Fordyce Lab The Fordyce Lab develops new microfluidic tools for making systems-scale, biophysical measurements of genomic interactions. |
 |
Frydman Lab The Frydman lab uses a multidisciplinary approach to address fundamental questions about molecular chaperones, protein folding and degradation. |
 |
Fuller Lab A major focus of our work concerns the mechanisms that regulate stem cell behavior. |
 |
Gitler Lab We investigate the mechanisms of human neurodegenerative diseases. |
 |
Greely Lab We work on ethical, legal, and social issues in the Biosciences, including genetics. |
 |
Greenleaf Lab Our lab focuses on developing methods to probe the genome and epigenome at the single-cell and single-molecule levels. |
 |
|
 |
Kay Lab We study gene/RNAi therapeutics and the mechanisms of non-coding RNA-induced gene regulation. |
 |
Kim Lab Research Areas: C. elegans aging, Human aging, automatic cell lineage analyzer, ModENCODE. |
 |
Kundaje Lab The Kundaje lab develops computational models of gene regulation by integrating diverse types of large scale functional genomic data. |
 |
Li Lab We are primarily interested in identifying and understanding sequence variations in the RNA and DNA. |
 |
Lipsick Lab Our laboratory studies the structure and function of chromosomes and chromatin in metazoans. |
 |
Montgomery Lab Our lab focuses on understanding the mechanisms by which genetic variation influences human traits. |
 |
Ormond Lab Master's Program in Human Genetics and Genetics Counseling. |
| Pringle Lab Applying the model-system approach to studies of yeast cell biology and the cellular and molecular biology of the cnidarian-dinoflagellate symbiosis. |
|
 |
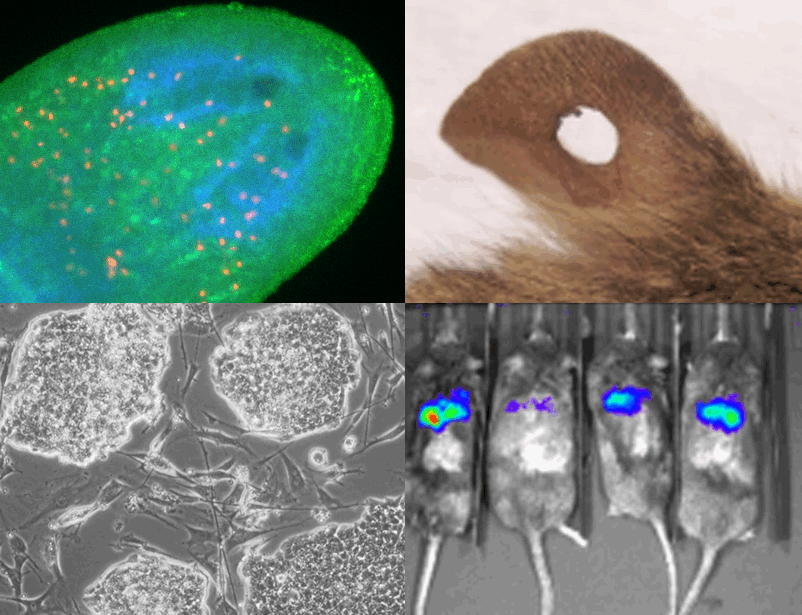
Pritchard Lab We are interested in a broad range of problems at the interface of genomics and evolutionary biology. |
 |
Sage Lab We investigate molecular and cellular mechanisms of tumorigenesis and regeneration, with a focus on stem cell biology. |
 |
Scott Lab Investigating how embryonic and later development is governed by proteins that control gene activity. |
| Sherlock Lab The Sherlock lab is a yeast genomics lab that uses both experimental and computational approaches. |
|
| Sidow Lab We have a diverse research program at the interface of computational and functional genomics. |
|
| Snyder Lab We are presently in an omics revolution in which genomes and other omes can be readily characterized. |
|
 |
Stearns Lab The central question behind our work is how the centrosome and primary cilium control cell function and influence development. |
| Steinmetz Lab The Steinmetz lab develops and applies interdisciplinary, genome-wide technologies to investigate the functions and mechanisms of genome regulation in health and disease. |
|
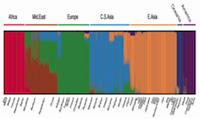
| Sun Lab My lab studies the molecular mechanism of transcription factors that govern the transformation of normal mammalian cells to a neoplastic state. |
|
 |
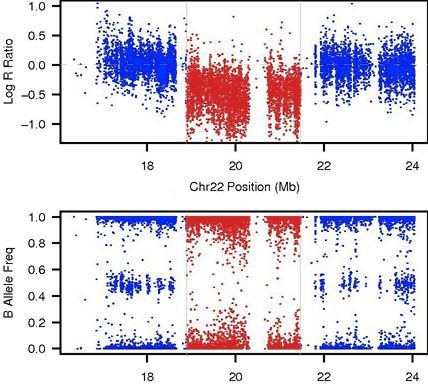
Tang Lab Research in our laboratory aims to uncover the evolutionary forces that have shaped the patterns of genetic variations. |
 |
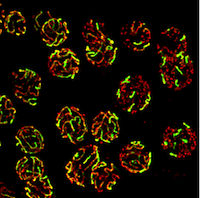
Urban Lab The Urban Lab investigates the effects of variation in human genomes on normal and abnormal brain development and function. |
 |
Villeneuve Lab Understanding the molecular and cellular mechanisms underlying the faithful inheritance and function of eukaryotic chromosomes. |
 |
Winslow Lab The goal of our lab is to understand the mechanisms of cancer progression and metastasis. |

